Smarter oncology second opinion — AI-powered, expert-reviewed, using your existing NGS report.
Read More

Cancer treatment decisions are becoming increasingly complex. Modern genomic testing can reveal several molecular alterations in a tumor, but understanding which therapy is most likely to work remains a challenge. At the AACR Annual Meeting 2026 (April 17–22, San Diego, USA), the Genomate Health research team will present new data demonstrating how computational reasoning can help interpret complex tumor genomic profiles and identify potentially effective treatments.
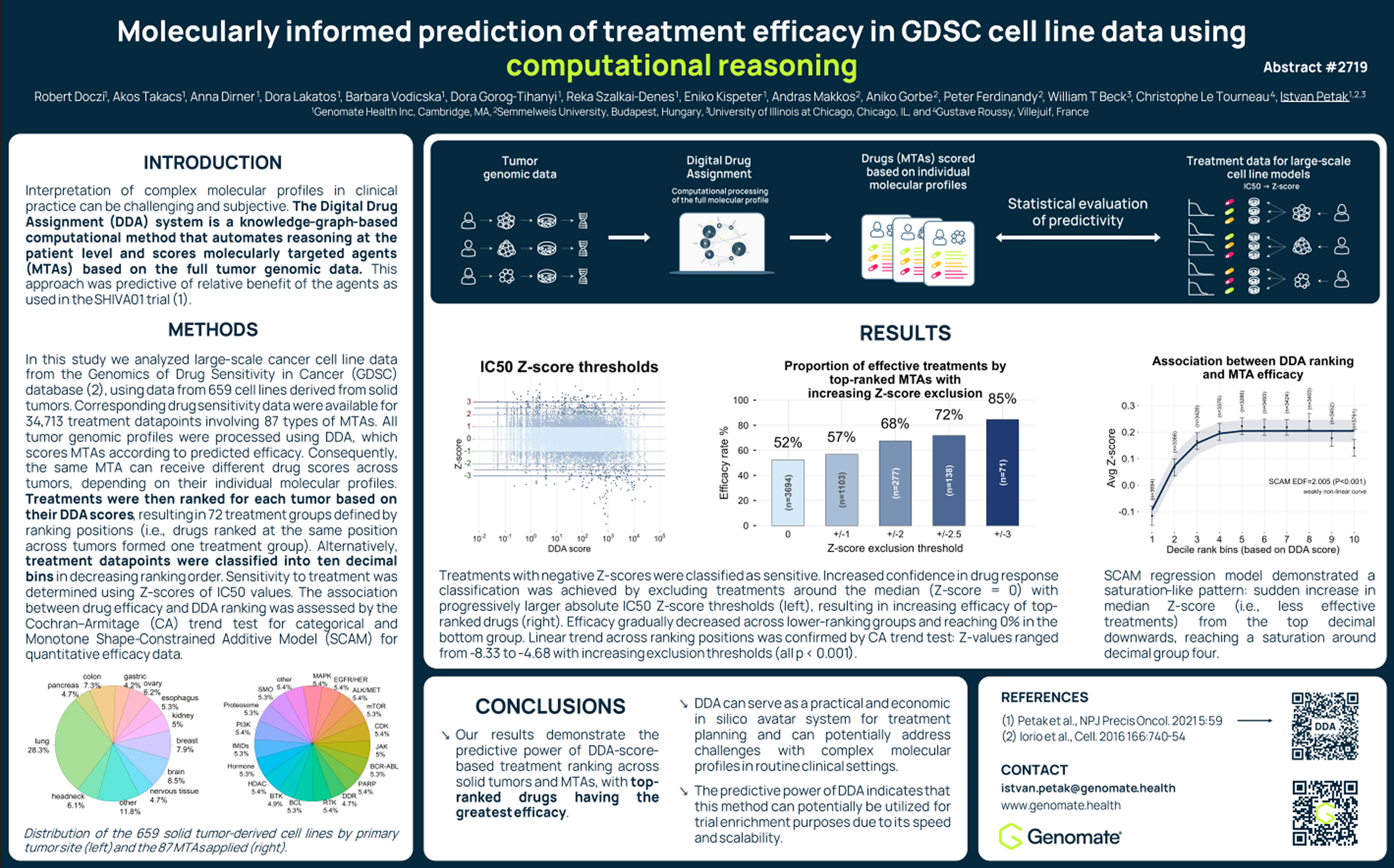
The study, titled “Molecularly informed prediction of treatment efficacy in GDSC cell line data using computational reasoning” evaluates the predictive performance of Genomate Health’s Digital Drug Assignment (DDA), commercially known as Genomate®, system using large-scale experimental drug sensitivity data.
Presenter: Istvan Petak, MD, PhD, Founder & Chief Scientific Officer, Genomate Health
Section: Integration of Clinical and Research Data
Poster Number: 2719
Date & Time: April 20, 2026, 2:00 PM - 5:00 PM
Digital Drug Assignment is a computational reasoning method that evaluates the relationship between tumor molecular alterations and potential cancer therapies. By integrating curated biological knowledge with tumor genomic data, the system ranks therapies according to their predicted effectiveness for each molecular profile.
To assess the predictive power of this approach, researchers analyzed data from the Genomics of Drug Sensitivity in Cancer (GDSC) database.
The analysis included:
Each tumor's genomic profile was analyzed using the DDA system, which assigned scores to therapies and ranked them by predicted efficacy.
Drug response was evaluated using IC50 Z-scores, a standard measure of drug sensitivity.
Among the therapies ranked highest by DDA:
Statistical analysis confirmed a highly significant trend across ranking groups (Cochran–Armitage trend test: Z = −10.42, p = 2.08 × 10⁻²⁵).
Increased confidence in the benchmark drug response classification was achieved by excluding treatments around the IC50 Z-score median with thresholds of progressively larger absolute values . Sensitivity among top-ranked treatments increased to 59%, 67%, 74%, and 83%, with increasing applied thresholds.
Across all thresholds, treatments in the lowest-ranked group consistently showed no sensitivity, reinforcing the predictive value of the ranking approach.
As genomic testing becomes more widely used in oncology, clinicians face an important challenge: translating complex genomic information into practical treatment decisions. A single tumor may contain dozens of molecular alterations, many of which interact with one another to influence treatment response. While next-generation sequencing can reveal these alterations, determining which therapy is most likely to work for a specific patient is often far from straightforward.
Computational reasoning approaches, such as Digital Drug Assignment, are designed to help address this challenge by systematically integrating genomic data with curated biological and clinical evidence. Instead of evaluating individual mutations in isolation, the system considers the full molecular context of a tumor and ranks potential therapies according to their predicted effectiveness.
The results of this study suggest that such approaches may help identify therapies with a higher likelihood of activity based on the molecular characteristics of each tumor. In the long term, tools that can interpret complex genomic profiles in this way may help physicians navigate large volumes of molecular data and translate sequencing results into more actionable treatment strategies.
More broadly, these results contribute to a growing body of evidence supporting the role of computational reasoning in precision oncology, an approach that aims to turn large volumes of molecular data into actionable insights for both clinical care and cancer research.
This research will be presented at the AACR Annual Meeting 2026 as a regular poster. It is part of a broader set of studies from the Genomate Health research team exploring how computational reasoning can support precision oncology. Two additional abstracts from our team will also be presented at the conference:
Abstract title: Molecularly-informed prediction of treatment efficacy in the GENIE BPC NSCLC cohort using computational reasoning ↗
Late-breaking abstract: Clinical efficacy of computational reasoning for personalized treatment planning in a pan–cancer cohort discussed by a French multidisciplinary tumour board: a real-world experience-based analysis ↗
If you are attending AACR 2026, we invite you to visit our poster presentations and meet the Genomate Health team to learn more about our research. Schedule a meeting with us during the conference to discuss the research and explore potential collaborations.