Smarter oncology second opinion — AI-powered, expert-reviewed, using your existing NGS report.
Read More

Genomic testing is now widely used to guide treatment decisions in non-small cell lung cancer (NSCLC). However, interpreting complex genomic profiles and selecting the most effective therapy remains a major challenge. At the AACR Annual Meeting 2026, researchers from Genomate Health will present new real-world evidence showing how computational reasoning can help predict treatment outcomes using large clinico-genomic datasets.
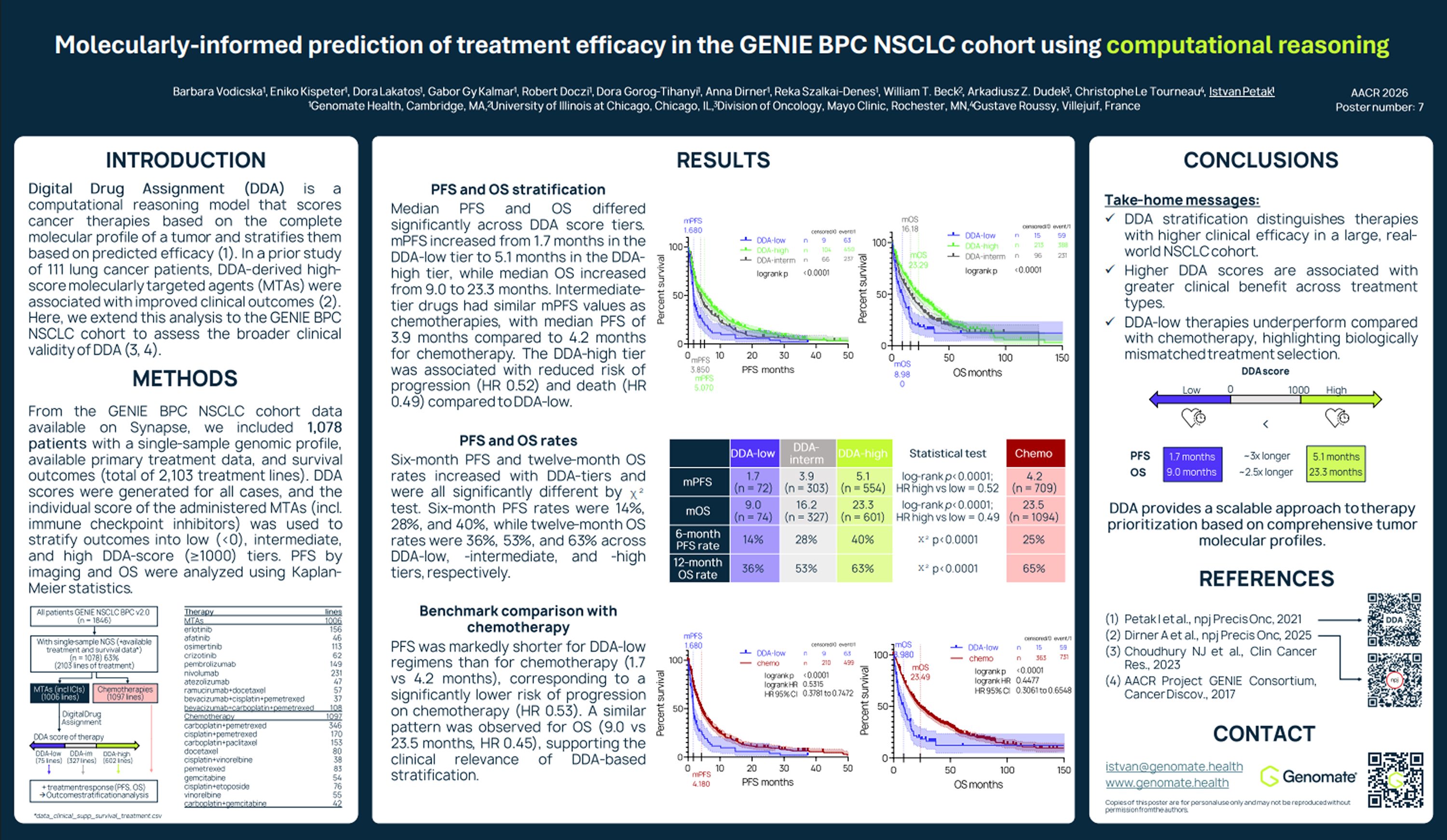
The study, titled “Molecularly-Informed Prediction of Treatment Efficacy in the GENIE BPC NSCLC Cohort using computational reasoning,” analyzes data from the AACR Project GENIE Biopharma Collaborative (BPC) cohort.
Presenter: Barbara Vodicska, PhD, Head of Translational Science, Genomate Health
Section: AACR Project GENIE: Predictive Models and AI
Poster Number: 7
Date & Time: April 19, 2026, 2:00 PM - 5:00 PM
The analysis examined the performance of Digital Drug Assignment (DDA), commercially known as Genomate®, a computational reasoning system that evaluates cancer therapies based on a tumor's full molecular profile. Rather than focusing on a single mutation, the system analyzes multiple genomic alterations simultaneously and ranks therapies according to their predicted effectiveness.
To assess how well this approach reflects real-world patient outcomes, the researchers analyzed data from 1,078 patients with NSCLC, representing 2,103 lines of treatment and including targeted therapies, immunotherapies, and chemotherapy. The dataset also included genomic sequencing results and survival outcomes for each patient. For every treatment administered, the system generated a DDA score based on the patient’s tumor profile. These scores were then used to group therapies according to their predicted level of effectiveness.
The analysis revealed a clear relationship between predicted treatment scores and clinical outcomes.
Patients receiving therapies with higher DDA scores experienced longer progression-free survival, meaning their cancer remained under control for a longer period. Median progression-free survival increased from 1.7 months for low-scoring therapies to 3.9 months for intermediate-scoring therapies and 5.1 months for high-scoring therapies.
Overall survival showed a similar pattern. Patients treated with higher-scoring therapies lived significantly longer, with median survival increasing from 9 months in the lowest score group to 23.3 months among those receiving the highest-scoring treatments. These differences were statistically significant.
Short-term survival indicators also improved with higher DDA scores. Both six-month progression-free survival rates and twelve-month overall survival rates increased consistently across the score groups.
The findings suggest that computational reasoning may help interpret complex genomic data and identify therapies that have a higher likelihood of benefiting individual patients. In lung cancer, genomic sequencing is increasingly used to guide treatment selection, particularly as targeted therapies and immunotherapies continue to expand. However, tumors often contain multiple genomic alterations, and the biological significance of these alterations can vary depending on their combination and context.
As a result, interpreting sequencing results and selecting the most appropriate therapy can be challenging, especially when multiple treatment options are available.
Computational reasoning systems such as Digital Drug Assignment (Genomate®) aim to address this challenge by integrating genomic data with curated biomedical knowledge, including evidence about molecular pathways, drug mechanisms, and previously reported treatment responses. By analyzing these relationships systematically, the system can rank therapies according to their predicted relevance for a specific tumor profile. Approaches like this may help clinicians make more informed treatment decisions by highlighting therapies that are most strongly supported by the available molecular evidence. As genomic testing becomes more widely adopted in oncology, tools that help translate sequencing results into practical treatment guidance may play an increasingly important role in precision cancer care.
This research will be presented at the AACR Annual Meeting 2026 as a regular abstract poster. It is part of a broader set of studies from the Genomate Health research team exploring how computational reasoning can support precision oncology. Two additional abstracts from our team will also be presented at the conference:
Regular abstract: Molecularly Informed Prediction of Treatment Efficacy in GDSC Cell Line Data Using Computational Reasoning ↗
Late-breaking abstract: Clinical efficacy of computational reasoning for personalized treatment planning in a pan–cancer cohort discussed by a French multidisciplinary tumour board: a real-world experience-based analysis ↗
If you are attending AACR 2026, we invite you to visit our poster presentations and meet the Genomate Health team to learn more about our research. Schedule a meeting with us during the conference to discuss the research and explore potential collaborations.